|
 |
- Search
| Obstet Gynecol Sci > Volume 58(1); 2015 > Article |
Abstract
Objective
To evaluate the type-specific human papillomavirus (HPV) persistence or regression in women with or less than low-grade cervical intraepithelial neoplasia (CIN).
Methods
This prospective cohort study included patients with or less than cytological low-grade squamous intraepithelial lesion (or histologically CIN 1 when biopsy was performed) combined with HPV infection. The cohort was collected from July 2006 to November 2011 at Korea University Guro Hospital. Follow-up was performed with liquid-based Papanicolaou test, hybrid capture 2 test, AnyplexTM II HPV 28 Detection, colposcopic biopsy if necessary every 4 months. All patients were prospectively observed without treatment.
Results
One hundred and thirty-seven patients were enrolled. Of these, 21 patients whose minimum follow-up periods were less than 8 months were excluded. Finally, one hundred sixteen patients were included and followed-up. Median follow-up period was 16 months. In case of high-risk HPVs, HPV 53 was the most prevalent type, followed by HPV 52, 68, 66, and 16. HPV 16 took 10.6 months to regress spontaneously, which was the longest period among the 10 most prevalent high-risk HPV genotypes. In case of spontaneous regression, HPV clearance was always accompanied by lesion clearance. A total of 13 patients showed disease progression either cytologically or histologically. Two cases of CIN 3 were confirmed by colposcopy-directed biopsy during follow-up, which were subsequently managed by conization.
Cervical cancer is the second most common cancer in women worldwide, and 80% of cases occur in developing countries [1]. Certain types of human papillomavirus (HPV) are the principal cause of invasive cervical cancer and cervical intraepithelial neoplasia [1]. More than 80 HPV types have been identified and subdivided into low-risk types, which are found mainly in genital warts, and high-risk types, which are frequently associated with invasive cervical cancer [2,3,4]. Most HPV infections are self-limiting and spontaneously regress within 1 to 2 years as a result of cell-mediated immunity. In one study, two-thirds of adolescents infected with low-risk HPV types spontaneously cleared their infections by 12 months, as did over 50% of those infected with high-risk HPV types [5]. In another follow-up study of adolescents and young women with low-grade squamous intraepithelial lesion (LSIL), 91% of HPV-infected individuals cleared their infections after 36 months of follow-up and only 5% to 10% of HPV infections progressed to persistent infection [6]. Persistent infection with certain HPV genotypes is an important factor in the development of cervical cancer. HPV 16 and HPV 18 are the most prevalent types and account for more than 70% of all invasive cervical cancers [7]. Infection with HPV 16 or HPV 18 tends to persist longer than infection with other HPV types [8], and persistence of infection with a high-risk HPV is a requirement for the development of cervical neoplasms such as cervical intraepithelial neoplasia (CIN) 2/3 and invasive cervical cancer. There is a marked difference between the HPV types associated with CIN grades 1, 2, or 3, and the relationship between the natural history of HPV and low-grade CIN has not been fully elucidated. In addition, previous meta-analyses reported a significant regional variation in overall prevalence and type distribution of HPV [9]. Although the most prevalent type worldwide is HPV 16 followed by HPV 18, in Asia HPV 58 is the most prevalent type [10]. In this study, we report the regression or persistence of specific human papilloma-virus genotypes in Korean women with low-grade CIN.
We evaluated data and clinical samples of patients diagnosed with cytologic LSIL or histologic CIN1 on biopsy combined with HPV infection at Guro Hospital College of Medicine at Korea University between July 2006 and November 2011. Women with cytologic evidence of high-grade squamous intraepithelial lesion (HSIL) or malignancy were excluded from this study group. All patients were prospectively observed without treatment and epidemiologic data concerning social and reproductive factors were obtained by private interviews with the patients. Korea University Guro Hospital institutional review board approved this study. All participants signed informed consent forms according to the recommendations of our institutional review board. Follow-up sampling was performed every 4 months with Papanicolaou (Pap) smear, HPV test, colposcopic biopsy or excisional biopsy, if necessary. In addition, data on duration of persistence or time to regression and corresponding disease status was obtained.
Enrolled patients were all tested with liquid-based Pap test, hybrid capture 2 test, and Anyplex II HPV 28 Detection system (Seegene, Seoul, Korea). The hybrid capture 2 test was conducted according to the manufacturer's instructions. This is a signal-amplified hybridization antibody capture assay using a luminometer with a specific probe mixture for high-risk HPV types 16, 18, 33, 35, 39, 45, 51, 52, 56, 58, 59, and 68 [11]. DNA-RNA hybrid products were captured on the surface of a microtiter plate coated with anti-DNA-RNA hybrids and the immobilized hybrids were reacted with alkaline phosphate-conjugated antihybrid monoclonal antibody. Light intensity was measured with a luminometer. The relative light unit for the recommended positive threshold of 1 pg/mL was used as a cutoff, and when the relative light unit was less than the cut-off the sample was considered to be negative.
Anyplex II HPV 28 was performed as recommended by the manufacturer using 5 µL DNA in each of the two 20-µL solutions with primer sets A or B [12]. This system uses HPV-specific double priming oligonucleotides for real-time polymerase chain reaction. In addition, it uses a primary HPV-specific probe (pitcher) carrying an artificial 5' extension and a secondary fluorescent tagging probe (catcher) specific to the primary probe. The catcher is activated by release of the 5' extension from the bound pitcher by the exonucleolytic activity of the polymerase. The 5'extension of the catcher binds to the catcher 3' and synthesize DNA and release quenching and emission of fluorescence. The catchers are tagged with a common fluorochrome for a set of three genotypes. Each catcher can be distinguished from others with the same fluorochrome by melting temperature during real-time polymerase chain reaction because the HPV types are clearly defined by their melting temperatures. This design constitutes the TOCETM system. In addition, analysis of the melting curve can be used to approximately quantify the viral load as low, intermediate, or high. In conclusion, Anyplex II HPV 28 is a fast, reliable test for the detection, differentiation, and quantification of multiple HPV genotypes that contribute to cervical cancer and sexually transmitted infection (HPV 6, 11, 16, 18, 26, 31, 33, 35, 39, 40, 42, 43, 44, 45, 51, 52, 53, 54, 56, 58, 59, 61, 66, 68, 69, 70, 73, and 82).
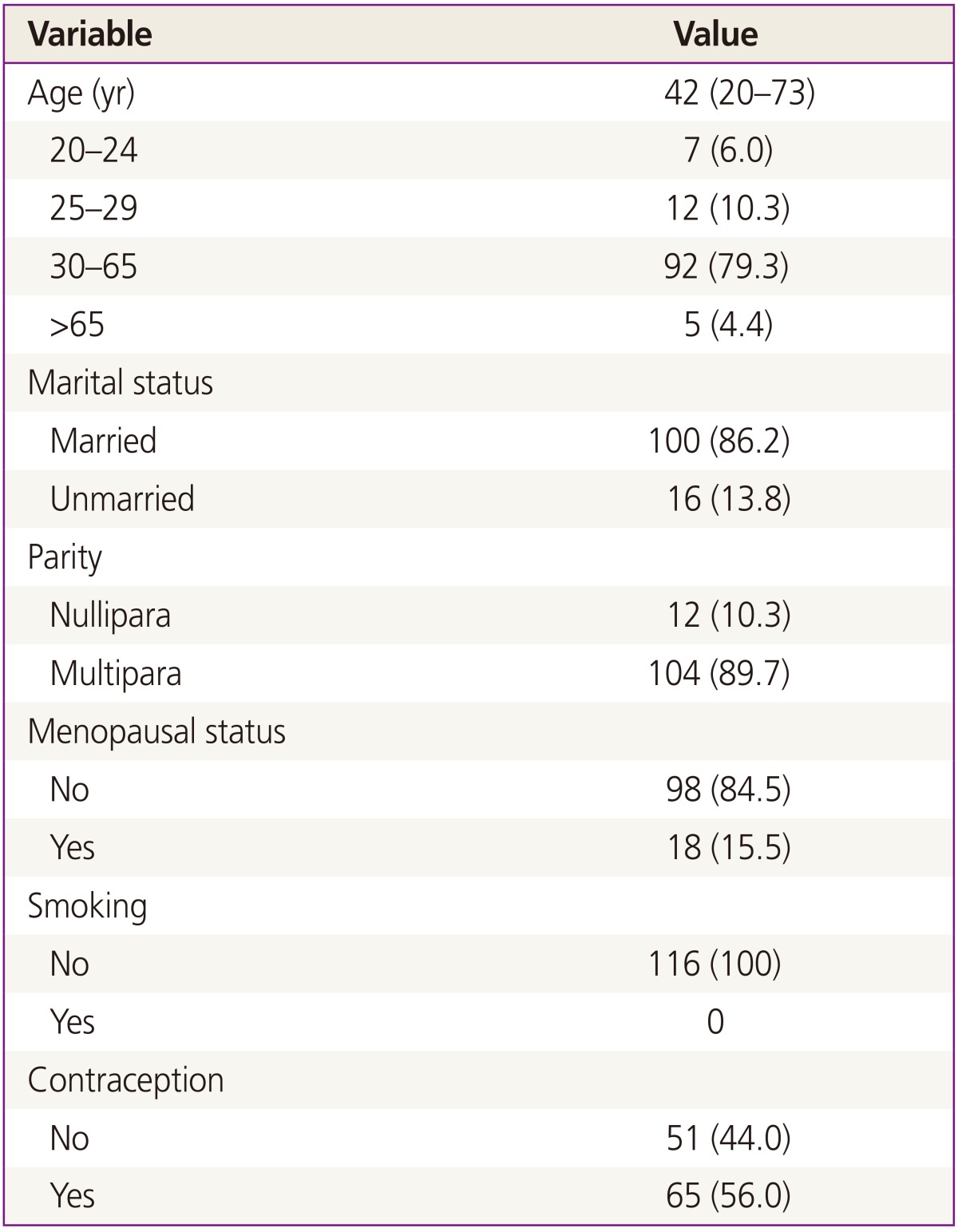
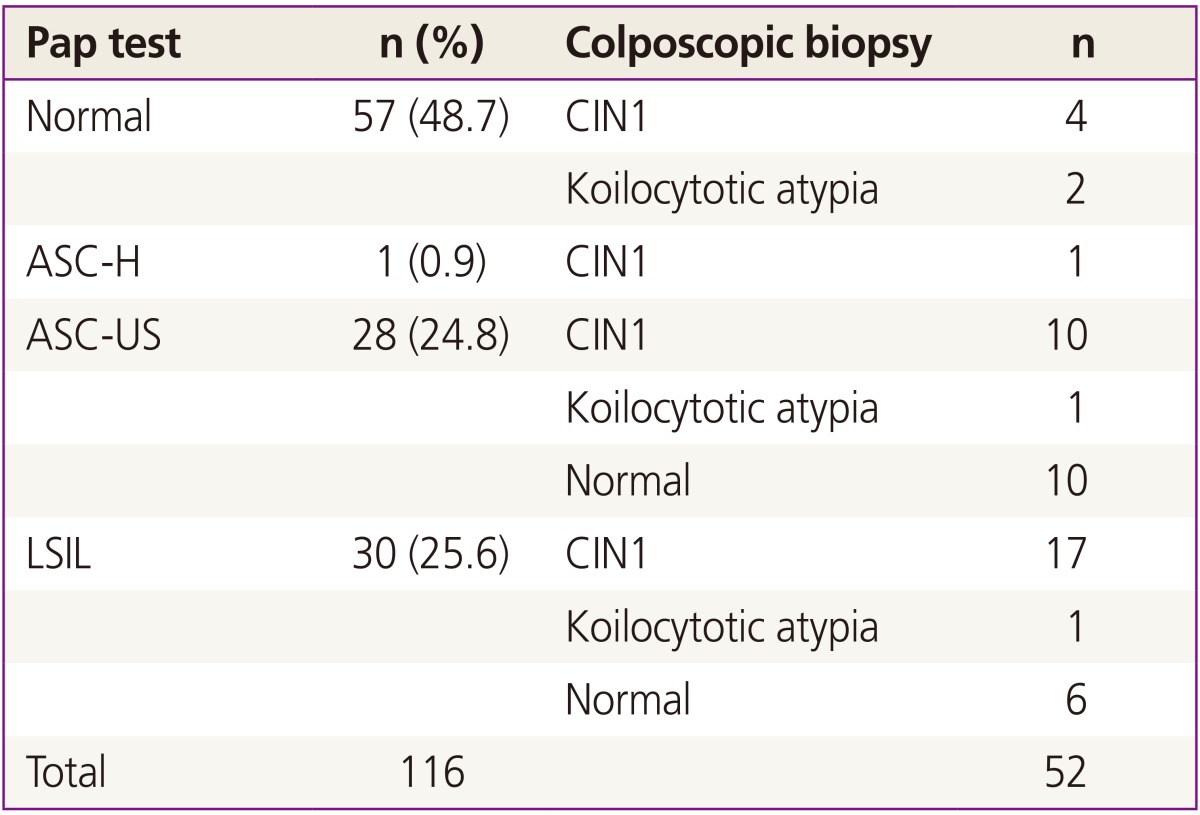
Among 137 patients who were initially enrolled, 21 patients with a minimum follow-up period of less than 8 months were excluded and a total of 116 patients were finally included and followed up in this study. Table 1 shows the demographic characteristics of patients. Median follow-up period was 16 months. Thirty patients were diagnosed with LSIL. Of them, biopsy was taken in 24 patients, including 6 normal, 17 CIN1, and 1 koilocytotic atypia. The remaining 6 did not take biopsies; 4 of them because of follow-up loss, 2 were taken only colposcopy. Individual Pap smear results and corresponding biopsy results were shown in Table 2.
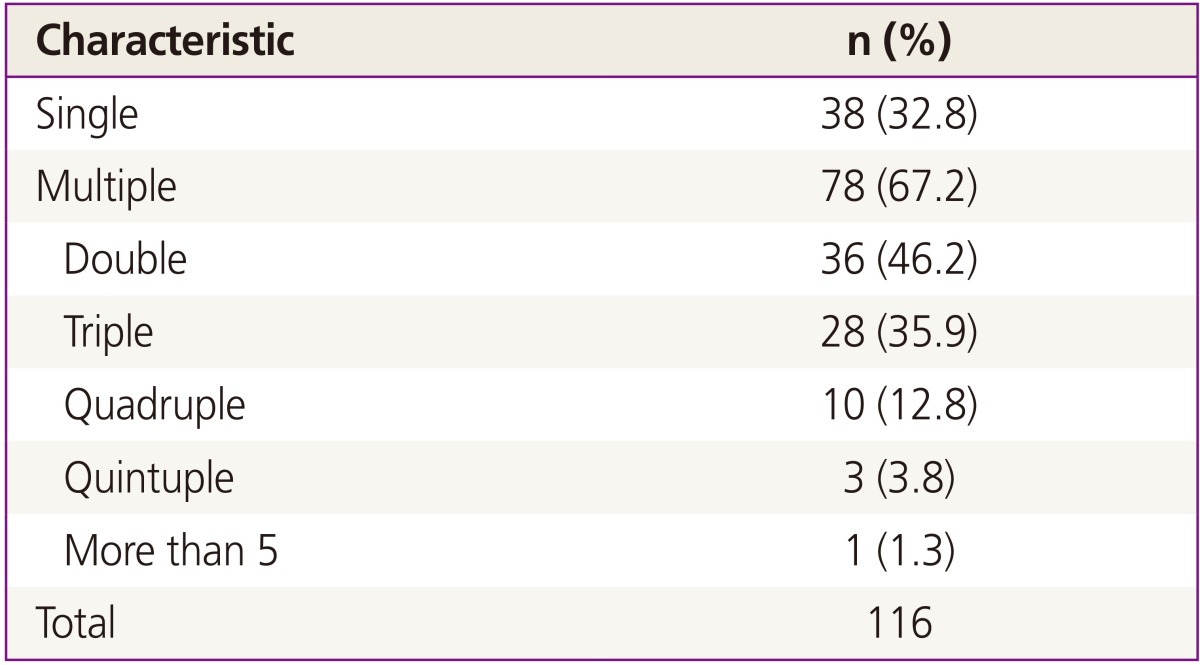
As shown in Table 3, multiple infections were common among patients in this study: 78 patients (67.2%) had infection with multiple HPV genotypes and only 38 women (32.8%) had a single type of HPV. Among women infected with multiple HPV types, double infection was the most common (n=36, 46.2%), followed by triple (n= 28, 35.9%), quadruple (n=10, 12.8%), and quintuple (n=3, 3.8%). Fig. 1 shows type-specific prevalence of HPV at enrollment including both high risk (HR) and low risk (LR) types. HPV 53 (n=33) was the most common type, followed by HPV 52 (n=25), 68 (n=22), 66 (n=20), and 16 (n=14). For LR HPVs, HPV 70 (n=27) was the most prevalent type.
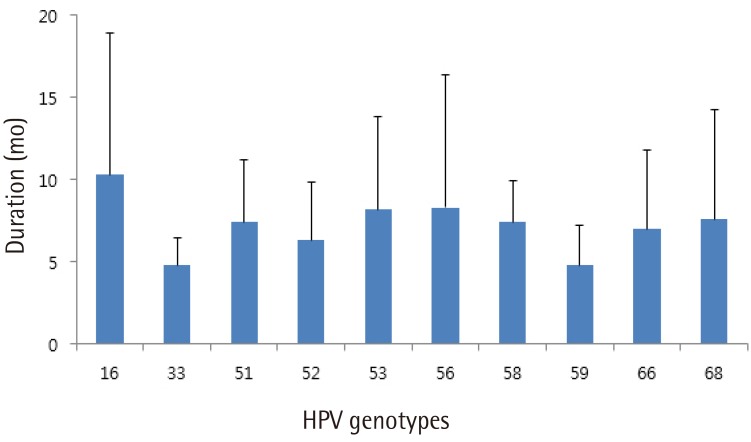
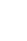
Fig. 2 shows the mean time to regression for the 10 most prevalent high-risk HPV genotypes. HPV 16 showed spontaneous regression at 10.3 months, which was the longest period to regression among the 10 high-risk HPV genotypes tested. In cases with spontaneous regression, HPV clearance was always accompanied by lesion clearance. A total of 13 patients showed disease progression either cytologically or histologically. Cytologically, 10 patients showed progression, including 6 patients from atypical squamous cells of undetermined significance to LSIL, 2 from normal to LSIL, 1 from normal to HSIL, and 1 from atypical squamous cells of undetermined significance to HSIL. Histologically, two patients were diagnosed with CIN3 and one patient with CIN2 during follow-up. Two of them underwent conization and the remaining one was lost to follow-up.
This study evaluated the prevalence and type-specific persistence of HPV in women with low-grade CIN in Korea. Although many previous studies have discussed the relationship between HPV infection and CIN or invasive cervical cancer, our understanding remains incomplete [3]. One limitation of these studies is the crudeness of the information. Because there are many variables that might affect the results, it is difficult to explain the geographic variation in cervical cancer incidence and the variability reported in different countries. Another limitation is that individual susceptibility to HPV was not considered in many studies [3]. There is a clear need for further research to understand the factors that determine whether a woman with HPV infection will clear the infection or become a persistent carrier. The results of this study provide a more complete understanding of the natural history of HPV infection.
HPV 53 was the most prevalent type in the present study, which is somewhat unusual compared with previous reports. An international meta-analysis showed that 70% of cases of invasive cervical cancer were associated with infection with HPV 16 (55%) or 18 (15%), and the next most common types were HPV 31, 33, 45, 52, and 58, which accounted for an additional 18% of cases. The most common HPV types in patients with HSIL were similar to those in patients with invasive cancer [13]. The prevalence of invasive cancer and HSIL is especially high in Asian patients with HPV 58 and 52 [14]. In a similar study of LSIL, HPV 16 was clearly the most prevalent HPV genotype worldwide and HPV 53 and 66, the so-called probable HR types, were relatively common in LSIL but were rarely found in invasive cancer and exhibited a squamous cell carcinoma/LSIL ratio lower than that for LR HPV type 6 [15]. In a large cohort study in Korea, the most prevalent HPV types were HPV 58, 16, 52, and 18, which together accounted for 73.9% of the HR HPV infections detected [16]. These discrepancies might be because the present cohort was collected at single medical center and has a relatively small sample size.
Although the most prevalent genotype in this study was HPV 53, HPV 16 was the most persistent HPV type. Together, HPV 16 and HPV 18 account for 35% of HPV-positive LSILs but nearly 70% of worldwide cervical cancers [2,14]. In prospective studies, HPV 16 infection was found to be more persistent and more likely to lead to progression to CIN3 than other HR HPV genotypes [17,18,19,20]. Although the degree of difference varies between genotypes, all HPV genotypes except HPV 16 and 18 were found to be unclearly relate to cervical cancers compared with LSIL [14]. In meta-analysis of Korean data, HPV 16, 58, and 18 were the three most common types in high-grade lesions, accounting for 62.0% of all high-grade lesions cases and 31.4% of low-grade lesions [16]. This result suggests that HPV genotyping might have a diagnostic role in separating LSIL cases with HPV 16 and 18 from LSIL cases with other HR genotypes.
Multiple HPV infections were common in this cohort, especially double infection (46.2%). According to previous studies, although there is no concrete evidence that previous infection can modulate the risk of secondary infection with HPV, already infected women are more likely to get another HPV infection than those who are not infected [20]. One previous study reported that women with HPV 58 were seven times more likely to get HPV 16 or 18 infections than uninfected women [21].
This is a prospective study designed to determine the natural history of HPV in Korea. The small sample size, the fact that it is not representative of the general Korean population, and the relative short duration of follow up are limitations of our study. However, our data suggest that distinction between HR HPV genotypes might explain the progression of LSIL and might be useful in the management of LSIL in clinical practice. The persistence of certain HR HPV types may explain the role of HPV in the pathogenesis of cervical cancer development from precancerous lesions. Further studies with a larger sample size and longer follow-up are needed to demonstrate whether persistence of specific HR HPV types is directly correlated with progression of low-grade lesions.
Acknowledgments
We are grateful for the work of HL Lee in data collection. We thank the women who participated in this study.
References
1. Parkin DM, Bray F, Ferlay J, Pisani P. Global cancer statistics, 2002. CA Cancer J Clin 2005;55:74-108. PMID: 15761078.


2. Muñoz N, Bosch FX, de Sanjose S, Herrero R, Castellsague X, Shah KV, et al. Epidemiologic classification of human papillomavirus types associated with cervical cancer. N Engl J Med 2003;348:518-527. PMID: 12571259.


3. Bosch FX, Lorincz A, Muñoz N, Meijer CJ, Shah KV. The causal relation between human papillomavirus and cervical cancer. J Clin Pathol 2002;55:244-265. PMID: 11919208.



4. Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, et al. Human papillomavirus is a necessary cause of invasive cervical cancer worldwide. J Pathol 1999;189:12-19. PMID: 10451482.


5. Brown DR, Shew ML, Qadadri B, Neptune N, Vargas M, Tu W, et al. A longitudinal study of genital human papillomavirus infection in a cohort of closely followed adolescent women. J Infect Dis 2005;191:182-192. PMID: 15609227.


6. Moscicki AB, Shiboski S, Hills NK, Powell KJ, Jay N, Hanson EN, et al. Regression of low-grade squamous intra-epithelial lesions in young women. Lancet 2004;364:1678-1683. PMID: 15530628.


7. Castellsagué X. Natural history and epidemiology of HPV infection and cervical cancer. Gynecol Oncol 2008;110(3 Suppl 2):S4-S7. PMID: 18760711.


8. Woodman CB, Collins S, Winter H, Bailey A, Ellis J, Prior P, et al. Natural history of cervical human papillomavirus infection in young women: a longitudinal cohort study. Lancet 2001;357:1831-1836. PMID: 11410191.


9. Bao YP, Li N, Smith JS, Qiao YL; ACCPAB members. Human papillomavirus type distribution in women from Asia: a meta-analysis. Int J Gynecol Cancer 2008;18:71-79. PMID: 17466054.

10. Bae JH, Lee SJ, Kim CJ, Hur SY, Park YG, Lee WC, et al. Human papillomavirus (HPV) type distribution in Korean women: a meta-analysis. J Microbiol Biotechnol 2008;18:788-794. PMID: 18467878.

11. Min KJ, So KA, Lee J, Hong HR, Hong JH, Lee JK, et al. Comparison of the Seeplex HPV4A ACE and the Cervista HPV assays for the detection of HPV in hybrid capture 2 positive media. J Gynecol Oncol 2012;23:5-10. PMID: 22355460.



12. Estrade C, Sahli R. Comparison of Seegene Anyplex II HPV28 with the PGMY-CHUV assay for human papillomavirus genotyping. J Clin Microbiol 2014;52:607-612. PMID: 24478495.



13. Smith JS, Lindsay L, Hoots B, Keys J, Franceschi S, Winer R, et al. Human papillomavirus type distribution in invasive cervical cancer and high-grade cervical lesions: a meta-analysis update. Int J Cancer 2007;121:621-632. PMID: 17405118.


14. Clifford GM, Smith JS, Aguado T, Franceschi S. Comparison of HPV type distribution in high-grade cervical lesions and cervical cancer: a meta-analysis. Br J Cancer 2003;89:101-105. PMID: 12838308.



15. Clifford GM, Rana RK, Franceschi S, Smith JS, Gough G, Pimenta JM. Human papillomavirus genotype distribution in low-grade cervical lesions: comparison by geographic region and with cervical cancer. Cancer Epidemiol Biomarkers Prev 2005;14:1157-1164. PMID: 15894666.


16. Kim MJ, Kim JJ, Kim S. Type-specific prevalence of high-risk human papillomavirus by cervical cytology and age: data from the health check-ups of 7,014 Korean women. Obstet Gynecol Sci 2013;56:110-120. PMID: 24327989.



17. Richardson H, Kelsall G, Tellier P, Voyer H, Abrahamowicz M, Ferenczy A, et al. The natural history of type-specific human papillomavirus infections in female university students. Cancer Epidemiol Biomarkers Prev 2003;12:485-490. PMID: 12814991.

18. Londesborough P, Ho L, Terry G, Cuzick J, Wheeler C, Singer A. Human papillomavirus genotype as a predictor of persistence and development of high-grade lesions in women with minor cervical abnormalities. Int J Cancer 1996;69:364-368. PMID: 8900368.


19. Molano M, Van den Brule A, Plummer M, Weiderpass E, Posso H, Arslan A, et al. Determinants of clearance of human papillomavirus infections in Colombian women with normal cytology: a population-based, 5-year follow-up study. Am J Epidemiol 2003;158:486-494. PMID: 12936904.


20. Woodman CB, Collins SI, Young LS. The natural history of cervical HPV infection: unresolved issues. Nat Rev Cancer 2007;7:11-22. PMID: 17186016.


21. Mendez F, Munoz N, Posso H, Molano M, Moreno V, van den Brule AJ, et al. Cervical coinfection with human papillomavirus (HPV) types and possible implications for the prevention of cervical cancer by HPV vaccines. J Infect Dis 2005;192:1158-1165. PMID: 16136457.


-
METRICS

-
- 21 Crossref
- 3,161 View
- 39 Download
- Related articles in Obstet Gynecol Sci